Diet is a fundamental regulator of host physiology, influencing metabolism, immunity and susceptibility to a wide range of diseases1,2,3,4,5. The gut microbiota has emerged as a key mediator of dietary effects, contributing to interindividual variation in the nature, direction and magnitude of responses to nutritional and molecular inputs13,14,15. Among the many physiological processes that are influenced by diet–microbiota interactions, regulation of adipose tissue remodelling has emerged as an intriguing yet poorly characterized area. Unlike brown adipose tissue (BAT), which is enriched in mitochondria and expresses thermogenic proteins, such as uncoupling protein 1 (UCP1), WAT stores excess energy. However, notably, WAT retains considerable plasticity and can acquire brown-fat-like properties under specific conditions, such as cold exposure and catecholamine stimulation. This process, termed browning or beiging, gives rise to cells characterized by enhanced mitochondrial biogenesis and expression of thermogenic and lipid-oxidation-related genes6,16,17,18. Several dietary interventions—including caloric restriction, ketogenic diets and deprivation of methionine, leucine or cysteine—have been reported to activate BAT and induce WAT browning19,20,21,22,23,24. The gut microbiota is thought to have a role in mediating these responses, potentially through the production of bioactive metabolites and immunomodulatory effects25,26,27,28,29,30. However, which members of the microbiota are involved in particular dietary contexts, and how diet–microbiota interactions are translated into signals that drive beige-fat biogenesis, remain poorly defined. In this study, we identify two orthogonal pathways that link low protein intake, specific microbial metabolic activity and host signalling networks to promote WAT browning.
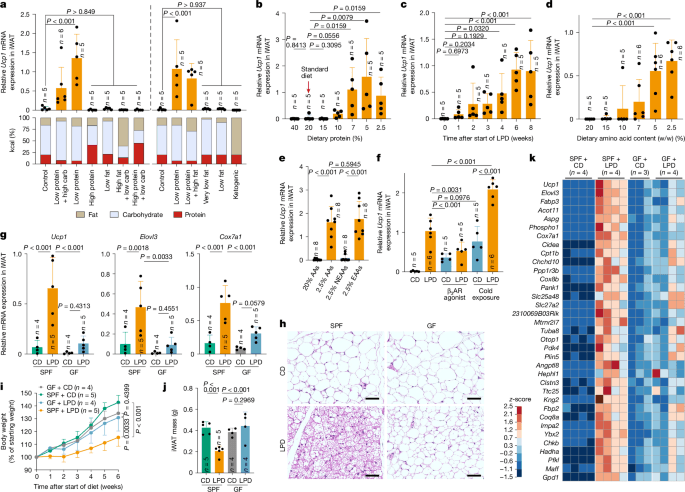
We first examined the effects of dietary modifications on WAT browning by formulating diets with varying proportions of protein, carbohydrate and fat. Although changes in fat content affected the total calorie count, all other diets were designed to be isocaloric. These diets were fed to specific-pathogen-free (SPF) C57BL/6 (B6) mice housed at room temperature (23 °C). Regardless of the lipid and carbohydrate content, feeding mice diets with low levels of protein significantly increased the expression of Ucp1 and Cox7a1 mRNA in the inguinal WAT (iWAT) (Ucp1 expression is shown in Fig. 1a, with Cox7a1 in Extended Data Fig. 1a. Throughout the manuscript, Ucp1 data are shown in the main figures and additional beige markers are provided mainly in the Extended Data). mRNA expression was assessed using three distinct methods31 with concordant results (Supplementary Fig. 1a). Feeding mice an LPD resulted in decreased iWAT mass (Supplementary Fig. 1b) and increased histological features of browning, including cells with multilocular lipid droplets, particularly in the central region adjacent to the inguinal lymph node (Extended Data Fig. 1b). B6 mice that were fed diets with a graded protein content showed a robust induction of Ucp1, Elovl3 and Cox7a1 in iWAT when dietary protein was reduced to 7% or less (a level approximately 60% lower than that of the control diet) (Fig. 1b and Extended Data Fig. 1c–e). Time-course analysis showed that these genes were induced within 2 weeks on a 7% protein diet and plateaued by weeks 6–8 (Fig. 1c, Extended Data Fig. 1f and Supplementary Fig. 1c). RNA sequencing (RNA-seq) analysis revealed that a broad range of beige-fat signature genes were upregulated in iWAT, including those involved in mitochondrial function (such as Cox7a1 and Cox8b), thermogenesis (such as Elovl3, Dio2 and Ucp1) and lipid metabolism (such as Cidea, Dio2, Cpt1b and Acott11); this upregulation began within 1 week of LPD feeding and became progressively more pronounced over the next 6 weeks (Extended Data Fig. 1g,h). The iWAT transcriptional profile induced by LPD feeding closely resembled that induced by a β3-adrenergic receptor agonist (Extended Data Fig. 1h). A reduction in dietary protein content also led to an increase in Ucp1, Elovl3 and Cox7a1 expression in interscapular BAT, although at lower magnitudes than that observed in iWAT (Extended Data Fig. 1i).
a, SPF B6 male mice were fed diets with varying proportions of protein, carbohydrate (carb) and fat for 6 weeks. Ucp1 mRNA expression in iWAT, normalized to Ppib, is shown together with each diet’s macronutrient composition. The left and right panels represent two independent experiments. b–e, Relative Ucp1 mRNA expression in iWAT in mice that were fed isocaloric diets containing the indicated protein concentrations for 6 weeks (b), a 7% LPD for the indicated durations (c), diets containing defined amino acids with varying total amino acid contents for 6 weeks (d) or diets containing 20% or 2.5% total amino acids (AAs), or with either EAAs or NEAAs reduced to 2.5%, for 6 weeks (e). f, SPF mice were fed a control diet (CD) or an LPD for 6 weeks and, during the final week, either received daily intraperitoneal injections of a β3-adrenergic receptor (β3AR) agonist (20 µg per mouse per day) or were exposed to cold (6 °C). g–k, SPF and GF mice were fed a CD or an LPD for 6 weeks. Ucp1, Elovl3 and Cox7a1 expression in iWAT (g), representative haematoxylin and eosin (H&E)-stained sections (h), body-weight change (i) and iWAT mass (j) are shown. Scale bars, 50 μm (h). k, Heat map showing genes enriched in iWAT of SPF + LPD mice compared with SPF + CD, GF + CD and GF + LPD mice (reads per kilobase per million reads (RPKM) ≥ 200, fold change ≥ 4, adjusted P < 0.05 with Benjamini–Hochberg correction). Each circle in the bar graphs represents an individual mouse. Data are mean ± s.d. Statistical tests: one-way ANOVA with Benjamini–Hochberg correction (a,c–g,j), two-tailed Mann–Whitney test for each comparison (b) and two-way ANOVA with Benjamini–Hochberg correction (i).
Feeding B6 mice a 7% protein diet for 6 weeks reduced body weight and whole-body fat volume, and enhanced glucose tolerance (Extended Data Fig. 2a–e). Mice showed no overt changes in appearance or behaviour (Supplementary Video 1), and did not exhibit increased faecal energy content or impaired lipid absorption (Extended Data Fig. 2f,g). In addition, whole-body lean volume and gastrocnemius muscle mass were mostly preserved (Extended Data Fig. 2h–j and Supplementary Video 2). These results indicate that the 6-week, 7% protein diet was generally well-tolerated and that the observed upregulation of browning markers and metabolic effects are unlikely to be driven mainly by pathological malabsorption or severe muscle wasting. Accordingly, this LPD protocol was used for all subsequent experiments, unless otherwise noted.
LPD feeding robustly induced signature genes of beige fat, even under thermoneutral conditions (30 °C) (Extended Data Fig. 2k). The LPD-induced formation of beige adipocytes in iWAT was observed across several mouse strains, including BALB/c and ICR, and was therefore not specific to B6 mice (Supplementary Fig. 2a). However, female or aged B6 mice showed attenuated browning, compared with male or young mice, respectively (Extended Data Fig. 2l,m). In addition, perigonadal WAT did not exhibit an induction of beige marker genes (Supplementary Fig. 2b). After reversion to a regular diet, beige marker expression and beige-adipocyte-like morphology declined (Extended Data Fig. 2n), indicating that LPD-induced browning in iWAT is both inducible and reversible. Increased expression of Ucp1 and metabolic effects were also observed when mice were switched from a high-fat diet (HFD) either to an LPD (Extended Data Fig. 3a–e) or to a high-fat and low-protein diet (HFD to HF/LPD; Extended Data Fig. 3f–j), although the magnitude of this induction was attenuated with prolonged or greater fat exposure. Collectively, these results indicate that LPD-induced browning is robust but adipose-depot-specific, and modulated by age, sex and dietary context.
Because protein type and digestibility might influence outcomes, we next examined diets containing defined amino acids instead of natural proteins. Decreasing the dietary amino acid content led to a marked increase in beige marker expression in iWAT, with maximal induction, comparable to that observed with a 7% LPD, achieved at 5% or less amino acids (Fig. 1d and Extended Data Fig. 3k). We then formulated diets with decreased levels of essential amino acids (EAAs) or non-essential amino acids (NEAAs). Reducing EAAs recapitulated the induction of beige marker genes that was observed with total amino acid limitation, whereas restricting NEAAs had no discernible effect (Fig. 1e and Extended Data Fig. 3l). We next examined the effect of reducing each individual EAA in the diet. Reduction of isoleucine, leucine, phenylalanine, tryptophan, lysine, methionine or threonine resulted in modest and variable inductions of browning (Extended Data Fig. 3m). These results suggest that although restriction of individual EAAs has limited effects, concurrent restriction of several EAAs engages complementary pathways to drive a more robust browning response.
We next investigated the mechanistic underpinnings of LPD-mediated iWAT browning. Feeding mice an LPD robustly induced the expression of Ucp1, Elovl3 and Cox7a1, to levels comparable with those induced by cold exposure (6 °C for 7 days) or treatment with a β3-adrenergic receptor agonist (Fig. 1f and Supplementary Fig. 3a). Notably, combining an LPD with cold exposure resulted in an additive increase in the expression of beige marker genes, suggesting that distinct—although possibly overlapping—mechanisms of browning are involved. Although immune cells have been implicated in cold-induced browning32,33,34,35,36,37,38, LPD-induced browning was preserved across several immune-deficient mouse models, including those lacking αβ and γδ T cells, B cells, innate lymphocytes, type 1, 2 or 17 immune molecules, myeloid cells or lymphoid tissues (Extended Data Fig. 4a). We therefore focused on immune-independent mechanisms.
We next assessed the role of the gut microbiota in LPD-induced browning using germ-free (GF) mice. Compared with SPF mice, GF mice exhibited significantly reduced expression of Ucp1,Elovl3, and Cox7a1 and fewer histological indicators of iWAT browning after being fed an LPD (Fig. 1g,h). In addition, although the LPD suppressed weight gain and WAT mass in SPF mice, no such effect was seen in GF mice (Fig. 1i,j). Low-level induction of Ucp1 was occasionally observed in LPD-fed GF mice, suggesting that a microbiota-independent pathway exists, although this response was not robust (Supplementary Fig. 3b). RNA-seq of iWAT from LPD-fed GF mice revealed a reduction in the expression of several beige signature genes, compared with LPD-fed SPF mice, including Ucp1, Cidea, Elovl3, Cox7a1, Cox8b, Clstn3 and Acot11 (refs. 39,40,41) (Fig. 1k). Gene Ontology enrichment analysis highlighted the reduced expression of genes involved in mitochondrial biogenesis and lipid metabolism in LPD-fed GF versus SPF mice (Supplementary Fig. 3c). Similar trends were observed in mice that were fed a low-EAA diet (Supplementary Fig. 3d). Treating adult SPF mice with an antibiotic cocktail also led to a significant reduction in the expression of Ucp1, Elovl3 and Cox7a1 in iWAT (Supplementary Fig. 3e). These findings indicate that the microbiota is a key driver of LPD-mediated beige-adipocyte induction.
We next aimed to identify microbiota-derived molecules that might contribute to iWAT browning. Non-targeted liquid chromatography–mass spectrometry (LC–MS) analysis of ileal and plasma samples from SPF and GF mice that were fed a control diet or an LPD revealed that an LPD increased the plasma levels of unconjugated bile acids, including cholic acid (CA) and muricholic acid (MCA), in SPF mice (Extended Data Fig. 4a). Subsequent targeted LC–MS analysis confirmed the increased plasma levels of CA, αMCA, βMCA and chenodeoxycholic acid (CDCA), as well as 7α- and 7β-dehydroxylation products42, including 7-oxo-deoxycholic acid (7oxoDCA), ursocholic acid (UCA) and ursodeoxycholic acid (UDCA), in LPD-fed SPF mice compared with GF mice and control-diet-fed SPF mice (Fig. 2a and Extended Data Fig. 4b), although no such trends were seen in the intestinal contents (Supplementary Fig. 4a). These bile acids exhibited agonistic activity in an in vitro FXR reporter assay, indicating activation of this bile acid receptor (Supplementary Fig. 4b). Therefore, we next investigated whether FXR signalling was involved in browning. Mice deficient in FXR (encoded by Nr1h4) exhibited a significantly reduced induction of Ucp1 and Elovl3, fewer morphologically beige cells in iWAT and increased weight gain during LPD feeding, compared with wild-type controls (Fig. 2b, Extended Data Fig. 4c–e and Supplementary Fig. 4c). By contrast, deletion of another prominent bile acid receptor, TGR5 (Gpbar1−/−), did not affect LPD-induced browning (Fig. 2c, Extended Data Fig. 4d,e and Supplementary Fig. 4c). These findings indicate that microbiota-derived bile acids promote beige-adipocyte induction during LPD feeding at least in part through an FXR-dependent, TGR5-independent pathway.
a, Plasma bile acids in GF or SPF mice fed a CD or an LPD were quantified by LC–MS/MS. ND, not detected. b–d,g, SPF mice of the indicated genotypes were fed a CD or an LPD for 6 weeks, and iWAT Ucp1 expression, normalized to Ppib, was measured. e,f, Uniform manifold approximation and projection (UMAP) plots of iWAT snRNA-seq data for Ucp1 and Nr1h4 (e) and for Col1a1, Wnt2, Pdgfra and Dpp4 (f) from GF or SPF mice fed a CD or an LPD. Each dot represents a nucleus, and red indicates expression of the indicated gene, with colour intensity reflecting expression levels. Beige and ASPC clusters are outlined by blue dashed circles. h, Hepatic Fgf21 mRNA expression and plasma FGF21 levels in GF or SPF mice fed a CD or an LPD. i–k, Fgf21+/+ and Fgf21−/− littermates fed a CD or an LPD were analysed for iWAT gene expression (i), body-weight changes (j) and plasma bile acids (k). In k, n = 5 per group. l, Plasma FGF21 levels in Nr1h4−/− and Nr1h4+/− mice. m, iWAT Ucp1 mRNA expression in SPF mice of the indicated genotypes (Adrb1−/−Adrb2−/−Adrb3−/− (Adrb1/2/3−/−), Adrb1−/−Adrb2−/− (Adrb1/2−/−), Adrb2−/−Adrb3−/− (Adrb2/3−/−) or Adrb1−/−Adrb3−/− (Adrb1/3−/−)) after 6 weeks on a CD or an LPD. n, Representative whole-mount iWAT images stained for tyrosine hydroxylase and UCP1 with iDISCO processing; dashed lines indicate iWAT boundaries. Scale bar, 2 mm. o–q, Wild-type (WT) SPF and GF mice (o), or SPF mice of the indicated genotypes (p,q), were fed a CD or an LPD for 6 weeks and received a daily β3-adrenergic receptor agonist or phosphate-buffered saline (PBS) during the final week. Relative iWAT Ucp1 was measured. Circles represent individual mice; data are mean ± s.d. Box plots show median, interquartile range and range. Statistical analyses: one-way ANOVA with Benjamini–Hochberg correction (b–d,g–i,l,m,o–q), two-way ANOVA with Benjamini–Hochberg correction (j) and two-tailed Mann–Whitney test (a,k).
Given the broad expression of FXR in the intestine, liver and WAT43, we used tissue-specific Nr1h4-knockout mice to define the organ-specific contribution of FXR signalling to iWAT browning. Deleting FXR in intestinal epithelial cells (Nr1h4fl/fl;Vil1-cre) or hepatocytes (Nr1h4fl/fl;Alb-cre) did not affect LPD-induced Ucp1 expression. By contrast, adipocyte-specific deletion of FXR (Nr1h4fl/fl;Adipoq-cre) significantly reduced Ucp1 and Elovl3 expression and morphologically beige cells in iWAT (Fig. 2d, Extended Data Fig. 4e and Supplementary Fig. 4c), suggesting that FXR has a key role in adipose tissue. To identify FXR-expressing cell types in iWAT, we performed single-nucleus RNA-seq (snRNA-seq) of iWAT from GF and SPF mice fed a control diet or an LPD. A putative beige-cell cluster expressing Ucp1 and Elovl3 was detected almost exclusively in LPD-fed SPF mice (Fig. 2e and Extended Data Fig. 5a,b). Among bile acid receptors, FXR showed a restricted pattern of expression and was enriched in cells expressing markers of adipose stem and progenitor cells (ASPCs), including Wnt2 and Dpp4 (refs. 44,45,46) (Fig. 2e,f and Extended Data Fig. 5a,b). Consistently, conditional deletion of FXR in Dpp4+ cells (Nr1h4fl/fl;Dpp4-cre) significantly attenuated the induction of Ucp1 and Elovl3 after LPD feeding (Fig. 2g and Supplementary Fig. 4c). Pseudotime analysis further suggested a de novo differentiation trajectory towards beige cells, characterized by a sequential induction of mitochondrial lipid metabolism and thermogenic genes (Supplementary Fig. 4d–f).
Given the robust nature of LPD-induced browning, we hypothesized that multiple, complex pathways are involved in addition to the bile acid–FXR axis. Considering the central role of the liver in regulating metabolic responses to diet, we performed bulk RNA-seq and quantitative PCR (qPCR) on liver samples. In SPF but not in GF mice, LPD feeding for 1 or 6 weeks consistently induced a number of genes, including Phgdh, (Psat1, Aldh1l2 and Asns (Extended Data Fig. 6a,b). As these genes function in serine biosynthesis, mitochondrial one-carbon metabolism and ammonia-dependent asparagine production47,48, their induction implicates the gut microbiota in linking dietary protein restriction to hepatic nitrogen assimilation and recycling. These genes are known to be regulated by activating transcription factor 4 (ATF4)49,50, and, consistently, other ATF4-inducible genes (Mthfd2, Nupr1, Chac1 and Gdf15) were similarly upregulated in a microbiota-dependent manner (Extended Data Fig. 6a,b). Notably, this transcriptional response was not induced by treatment with a β3-adrenergic receptor agonist (Extended Data Fig. 6b), highlighting the unique nature of LPD-induced microbiota-dependent signalling in the liver.
Notably, Fgf21, another ATF4-inducible gene51, was markedly upregulated in the livers of SPF mice, but not GF mice, as early as 1 week after LPD initiation, with corresponding increases in circulating FGF21 (Fig. 2h and Extended Data Fig. 6b). Fgf21−/− mice phenocopied GF mice, showing a minimal induction of beige marker genes in iWAT and no suppression of weight gain during LPD feeding (Fig. 2i,j). However, administering recombinant FGF21 was insufficient to induce iWAT browning in control-diet-fed GF mice (Supplementary Fig. 5a), indicating that additional microbiota-dependent signals, including the bile acid–FXR axis, are required. Indeed, the gene-expression profiles of iWAT from Fgf21−/− and Nr1h4−/− mice showed distinct patterns (Supplementary Fig. 5b), and Nr1h4 and Klb (encoding a crucial component of the FGF21 receptor) were expressed in distinct cell populations within iWAT (Extended Data Fig. 5b). Moreover, LPD feeding resulted in comparable plasma bile acid profiles in Fgf21−/− and wild-type mice (Fig. 2k), and, conversely, plasma FGF21 levels were not significantly altered between Nr1h4−/− and wild-type mice (Fig. 2l). These findings suggest that FXR and FGF21 act through parallel, non-redundant pathways that collectively contribute to beige cell induction in iWAT.
To identify the hepatic cell types that express Fgf21, we performed snRNA-seq on liver tissue. Fgf21 and other ATF4-dependent genes were detected in hepatocyte clusters from LPD-fed SPF mice, but not in other liver cell types or in GF or control-diet-fed mice (Extended Data Fig. 7). A similar pattern of expression was observed for Cyp39a1 and Csad (Extended Data Fig. 7). Given the roles of CYP39A1 in non-canonical bile acid biosynthesis and CSAD in taurine synthesis52, these findings suggest that hepatocytes respond to LPD feeding by coordinately upregulating not only Fgf21, but also genes involved in bile acid production.
As catecholamine signalling is central to adipose browning6,16,17,40,53,54,55, we next assessed LPD-mediated beige-cell induction in mice deficient in β1- (Adrb1), β2- (Adrb2) or β3- (Adrb3) adrenergic receptors. Loss of β3-adrenergic signalling, either alone or combined with β1 or β2 deficiency, resulted in a severe impairment in beige-cell induction, whereas deletion of β1 and β2 had no effect (Fig. 2m and Extended Data Fig. 8a), suggesting that LPD-induced browning relies specifically on β3-adrenergic receptor signalling. Consistently, whole-mount immunostaining of iWAT revealed a pronounced remodelling of tyrosine hydroxylase-positive sympathetic neurons and vasculature in SPF but not in GF wild-type mice in response to an LPD (Fig. 2n and Extended Data Fig. 8b). Specifically, sympathetic neurons developed finer, denser networks, particularly in regions with a strong accumulation of UCP1-expressing cells. By contrast, LPD-fed Fgf21−/− and Nr1h4−/− SPF mice exhibited markedly reduced sympathetic innervation, resembling the pattern observed in GF mice (Fig. 2n and Extended Data Fig. 8c). Notably, administration of a β3-adrenergic receptor agonist increased Ucp1 and Cox7a1 expression in control-diet-fed GF mice as well as Nr1h4−/− and Fgf21−/− mice to levels comparable with those in LPD-fed SPF mice (Fig. 2o–q and Extended Data Fig. 8d–f). These findings suggest that FXR and FGF21 signal through distinct pathways that converge on the promotion of sympathetic innervation and β3-adrenergic signalling during LPD-induced browning.
Browning induced by 20 bacterial strains in mice
To assess whether LPD-induced iWAT browning is transmissible via the microbiota, we transplanted small-intestinal luminal contents from LPD-fed SPF donor mice (donors A and B) into GF mice (Fig. 3a). Browning was observed in recipient mice irrespective of the transplant source, but only when recipients were fed an LPD (Fig. 3b), indicating that ongoing diet–microbiota interactions are required. Similar results were obtained using faecal transplants (Supplementary Fig. 6a). To identify a bacterial community capable of mediating browning, we selected mouse B28, which exhibited the strongest induction of Ucp1 after receiving small-intestinal contents from LPD-fed mouse B and being maintained on an LPD (Fig. 3b and Supplementary Fig. 6b). Transferring small-intestinal contents from mouse B28 to new GF recipients recapitulated the robust iWAT browning upon LPD feeding (Fig. 3c and Supplementary Fig. 6c). We selected mouse B28-1, which exhibited the strongest beige adipocyte induction, for subsequent analyses (Fig. 3c). From the small-intestinal contents of mouse B28-1, we isolated 18 bacterial strains (mu18-mix) (Fig. 3d). However, colonizing GF mice with the mu18-mix was insufficient to induce browning (Fig. 3e). Adding two further isolates, Blautia sp. (St.27G3) and Turicibacter sp. (St.80E1), yielded a 20-strain consortium (mu20-mix) that robustly induced browning after LPD feeding, reaching levels comparable to those of SPF mice (Fig. 3f,g), indicating that these two strains are required for the full beige-inducing activity of the consortium. GF mice inoculated with the mu20-mix and fed an LPD showed increased plasma levels of CA, CDCA, 7oxoDCA, UCA and UDCA (Fig. 3h), as well as FGF21 (Fig. 3i). By contrast, Nr1h4−/− and Fgf21−/− GF mice inoculated with the mu20-mix exhibited markedly reduced beige-cell induction (Fig. 3j), further supporting the requirement for FXR and FGF21 signalling in LPD-induced browning mediated by defined members of the microbiota.
a, Schematic illustrating the strategy to isolate browning-inducing commensal strains from the gut microbiota of SPF mice. SI, small intestine. b,c,e,f, GF mice were colonized with ileal microbiota from LPD-fed SPF mouse A, SPF mouse B (b) or exGF mouse B28 (c), or with defined bacterial consortia (mu18-mix in e or mu20-mix in f). Mice were then fed an LPD for 6 weeks, and relative iWAT Ucp1 expression was measured by qPCR. d, The ileal microbiota compositions of mouse B, mouse B28 and mouse B28-1 were determined by 16S rRNA gene sequencing. The 20 strains isolated from mouse B28-1 are listed. g, Representative H&E-stained iWAT sections from the indicated groups. Scale bars, 50 μm. h,i, Plasma bile acid (h) and FGF21 (i) concentrations in the indicated mice were quantified by LC–MS/MS and by enzyme-linked immunosorbent assay (ELISA), respectively. j, iWAT Ucp1 expression in GF Nr1h4−/− or GF Fgf21−/− mice inoculated with mu20-mix or vehicle control and fed a CD or an LPD for 6 weeks. k, GF mice were colonized with the indicated mouse-derived bacterial consortia (mu20-mix, mu3-mix, mu5-mix or mu7-mix) and fed a CD or an LPD for 6 weeks. iWAT Ucp1 expression and hepatic Fgf21 expression were measured. Circles represent individual mice; data are mean ± s.d. Box plots show median, interquartile range and range. Statistics: one-way ANOVA with Benjamini–Hochberg correction for multiple comparisons (b,c,e,f,i–k) or two-tailed Mann–Whitney test for each comparison (h).
Having confirmed that a small microbial community is sufficient to promote LPD-induced browning, we next sought to identify human-associated microorganisms that are capable of exerting this activity. To this end, we recruited 25 healthy volunteers and performed 18F-fluorodeoxyglucose positron emission tomography (FDG-PET), which detects beige or brown adipose depots9,10,11,12. Approximately 40% of volunteers exhibited supraclavicular FDG accumulation (Fig. 4a). We transplanted faecal samples from the top four FDG-positive volunteers (T17, T10, T19 and T18) into GF mice and placed the mice on either a control diet or an LPD (Extended Data Fig. 9a). In exGF mice colonized with microbiota from donors T17, T10 or T19, LPD feeding robustly induced Ucp1 expression and morphologically beige adipocytes in iWAT (Fig. 4b and Extended Data Fig. 9b). Faecal microbiota transplantation from donor T18 was much less effective, probably due to unsuccessful engraftment of key effector strains. We also transplanted GF mice with faecal samples from donors FF2 and T07, who exhibited intermediate or no FDG accumulation, respectively. Mirroring the human phenotype, FF2 recipient mice exhibited an intermediate degree of browning, whereas T07 recipients showed no evidence of browning (Fig. 4c and Extended Data Fig. 9c).
a, Standardized uptake values (SUV) of FDG in the supraclavicular regions of each volunteer. Columns highlighted in red and blue indicate samples that were selected for follow-up analysis, and representative FDG-PET images are shown as inserts. b–f, GF B6 mice were colonized with faecal microbiota from the indicated individuals (b,c) or with defined bacterial consortia (d) and then fed either a CD or an LPD for 6 weeks. Relative iWAT Ucp1 expression, normalized to Ppib, was determined by qPCR (b–d). Plasma bile acids (e) and FGF21 (f) were quantified by LC–MS/MS and ELISA, respectively. g, iWAT Ucp1 expression in GF Nr1h4−/− and Fgf21−/− mice colonized with the T19-derived 33-mix and fed a CD or an LPD. h, List of the 33 T19-derived strains, as determined by 16S rRNA sequencing. i–m, GF B6 mice were colonized with the indicated bacterial consortia selected from the 33 T19-derived strains and fed a CD or an LPD for 4 weeks. In j, the presence (+) or absence (−) of each bacterial group in the consortium is indicated. Expression of Ucp1 in iWAT (i-k) and Fgf21 in the liver (l), normalized to Ppib, was measured by qPCR. m, Plasma bile acid concentrations in LPD-fed gnotobiotic mice were quantified by LC–MS/MS (n = 8 per group). Circles represent individual mice; bar heights indicate exact SUV values (a) or mean ± s.d. (b–d,f,g,i–l). Box plots show median, interquartile range and data range. Statistics: one-way ANOVA with Benjamini–Hochberg correction for multiple comparisons (b–d,f,g,i–l) or two-tailed Mann–Whitney test for each comparison (e,m).
Next, we isolated 33 strains from faecal samples from T10 microbiota-recipient mice (T10-4 and T10-5) and another 33 strains from T19 microbiota-recipient mice (T19-5 and T19-6) (Fig. 4b,h and Extended Data Fig. 9d). GF mice were colonized with each of these 33-strain mixtures. Mice that were colonized with the T19-derived 33-strain mixture exhibited robust beige-cell induction after LPD feeding, whereas those colonized with the T10-derived mixture showed substantially less induction (Fig. 4d and Extended Data Fig. 9e). Mice that were colonized with the T19-derived 33-mix exhibited increased plasma levels of CA, 7oxoDCA, UCA, CDCA and UDCA, as well as FGF21 (Fig. 4e,f). By contrast, Nr1h4−/− or Fgf21−/− GF mice inoculated with the T19-derived 33-mix failed to significantly upregulate Ucp1 in iWAT (Fig. 4g), indicating that these strains promote beige-cell induction through activation of both the FXR and the FGF21 pathway, similar to the mu20-mix.
To identify a minimal effector consortium, we divided the 33 T19-derived strains into 2 phylogenetic groups: a 19-strain group comprising Bacteroides, Enterococcus, Erysipelotrichaceae and other phyla (19 BEEO), and a 14-strain group comprising Ruminococcaceae and Lachnospiraceae (14 RL) (Fig. 4h). Gnotobiotic mice colonized with the 19 BEEO-mix recapitulated the robust induction of Ucp1 in iWAT that was observed with the parental 33-mix, whereas mice colonized with the 14 RL-mix did not (Fig. 4i). We further subdivided the 33 T19-derived strains into 5 phylogenetic groups (Fig. 4h) and assessed the effect of excluding each on iWAT browning. Excluding the five strains classified as other phyla (5 Others) led to the greatest reduction in Ucp1 induction (Fig. 4j). We thus compared the effects of colonization with the 5 Others-mix versus the remaining 14 strains from the 19 BEEO group (14 BEE). The 5 Others-mix efficiently induced beige marker genes in an LPD-dependent manner, whereas the 14 BEE strains produced only a modest effect (Fig. 4k and Extended Data Fig. 9f). Colonization with the 5 Others-mix, but not the 14 BEE-mix, also increased hepatic Fgf21 mRNA expression and plasma bile acid concentrations to levels comparable with those observed in mice colonized with the parental 19 BEEO-mix (Fig. 4l,m).
Additional dropout experiments were performed by inoculating GF mice with all possible four-strain permutations of the 5 Others-mix, each omitting one of the five strains. Notably, omitting Romboutsia timonensis (St.31) prevented engraftment of the remaining four strains and markedly reduced beige-cell induction (Fig. 5a and Extended Data Fig. 10a), suggesting that R. timonensis acts as a supporter strain that facilitates colonization by the others. Excluding Adlercreutzia equolifaciens (St.3), Eubacteriaceae sp. (St.4) or Bilophila sp. 4_1_30 (St.14) did not impair colonization by the remaining strains but substantially affected their ability to induce iWAT browning (Fig. 5a and Extended Data Fig. 10a). By contrast, omitting Parasutterella excrementihominis (St.29) had no effect on beige-cell induction (Fig. 5a). Therefore, whereas the Parasutterella strain is dispensable, all four remaining strains (Adlercreutzia, Eubacteriaceae, Bilophila and Romboutsia; hereafter referred to as ‘hu4’ strains) are essential for the observed LPD-mediated browning effect by human-associated strains.
a, GF B6 mice were colonized with four-strain permutations of the 5 Others-mix; included strains are indicated by + and the excluded strain by –. Mice were fed an LPD for 4 weeks, and iWAT Ucp1 expression, normalized to Ppib, was measured by qPCR. b, GF mice were inoculated with faecal microbiota from participant T07, fed a HFD (60% kcal fat, 20% protein) for 4 weeks, then switched to an LPD and given two oral doses of the hu4 strains or vehicle control (−). Longitudinal changes in body weight are shown. An oral glucose tolerance test (OGTT) was performed after 4 weeks on the LPD. After 6 weeks on the LPD, Ucp1 and Elovl3 expression in iWAT and BAT, hepatic Fgf21 expression, iWAT mass and plasma cholesterol, triglycerides and ALT were assessed. IU, international units. c, In vitro bile acid metabolic capacity of individual members of the 5 Others-mix after incubation with 50 µM tauro-CA. d, Metatranscriptomic analysis of bacterial RNA from the caecal contents of gnotobiotic mice colonized with the T19-derived 33-mix and fed a CD or an LPD. Reads mapped to the nrfA loci of Bilophila sp. 4_1_30 (St.14) and A. equolifaciens (St.3) are shown. e, NrfA homologues and predicted signal peptides in human isolates. eggNOG-mapper was used for annotation, and SignalP 6.0 was used to identify signal peptides. Percentage identity indicates similarity to the closest NrfA protein. E-value represents the expected value. Length denotes the aligned protein length. SP, standard signal peptide. Circles represent individual mice; data are mean ± s.d. (a,b) or exact bile acid concentrations (c). Statistics: one-way ANOVA with Benjamini–Hochberg correction (a), two-way ANOVA (b, line graphs) and two-tailed unpaired t-test (b, bar graphs).
To assess the metabolic effect of the hu4-strain consortium, GF mice were inoculated with faeces from participant T07, whose microbiota lacked browning capacity, and were fed a HFD. Mice were then treated with an LPD alone or combined with two oral doses of the hu4 strains. Whereas the LPD alone was somewhat efficacious, combination treatment with the hu4 strains led to significantly greater body-weight loss, increased expression of Ucp1 and Elovl3 in both iWAT and BAT, increased hepatic Fgf21 expression, reduced iWAT mass, decreased levels of plasma cholesterol, triglyceride and alanine transaminase (ALT) and improved glucose tolerance (Fig. 5b). These effects occurred with minimal reduction in muscle mass and no increase in plasma creatine kinase levels (Extended Data Fig. 10b), suggesting that WAT browning mediated by specific microbiota members enhances the metabolic effects of LPD feeding without overt muscle damage.
Microbial ammonia production induces hepatic FGF21
To elucidate how the hu4 strains promote LPD-induced browning, we examined their bile-acid-metabolizing capabilities. All four strains exhibited bile salt hydrolase activity, converting tauro-CA to CA in vitro (Fig. 5c). Furthermore, the supporter R. timonensis (St.31) strain encodes a putative 7α-hydroxysteroid dehydrogenase (7αHSDH) (Extended Data Fig. 10c) and uniquely converted tauro-CA into 7oxoDCA (Fig. 5c), which may facilitate colonization by the other strains and FXR activation in iWAT.
We next performed genome sequencing of the 33 T19-derived strains (Supplementary Table 1) and transcriptomic profiling of caecal microbiota from gnotobiotic mice colonized with this consortium and fed either a control diet or an LPD. LPD feeding selectively upregulated nitrogen-metabolism-related genes in the 19-BEEO and hu4 strains, but not in the 14-BEE or 14-RL strains (Extended Data Fig. 10d). Notably, expression of nrfA, which encodes and enzyme that catalyses the reduction of nitrite into ammonia, was greatly increased in Bilophila sp. 4_1_30 (St.14) and A. equolifaciens (St.3) upon LPD feeding (Fig. 5d). Unlike most nrfA homologues, the nrfA genes in these strains uniquely contain lipoprotein signal peptides (LSPs) (Fig. 5e), a feature that is rare in the human microbiome and mostly restricted to Desulfobacterota (Extended Data Fig. 10e and Supplementary Table 2). Such signal peptides are predicted to localize NrfA to the inner membrane, enabling periplasmic dissimilatory nitrate reduction to ammonium (Fig. 6a). NrfA uses formate as an electron donor through its partner enzyme, formate dehydrogenase (FDH)56. FDH contains a molybdopterin cofactor in its active site, which can be inhibited by treatment with tungsten57. Adding tungsten to the drinking water significantly suppressed both iWAT browning and plasma FGF21 levels in mice colonized with the 19 BEEO-mix, without altering strain engraftment or plasma bile acid levels (Fig. 6b–d and Extended Data Fig. 11a).
a, Schematic illustrating nitrite reduction by NrfA in conjunction with formate dehydrogenase (FDH). Mo, molybdopterin. b–d, GF B6 mice were colonized with 19 BEEO-mix and fed a CD or an LPD for 3 weeks. During the final 2 weeks, mice received 0.1% or 0.5% sodium tungstate dihydrate (W) in the drinking water. Relative iWAT Ucp1 expression (b), plasma FGF21 levels (c) and plasma bile acid concentrations (d) were assessed. In d, n = 8 per group. e, GF mice colonized with two strains selected from the hu4-mix (included, +; excluded, –) were fed a CD or an LPD for 4 weeks, and iWAT Ucp1 expression was quantified. f,g, GF mice were colonized with ΔnrfA or wild-type Bilophila sp. 4_1_30 (St.14) together with Romboutsia timonensis (St.31) (Rombo) and fed a CD or an LPD for 4 weeks; iWAT Ucp1 (f) and hepatic Fgf21 (g) expression were measured by qPCR. h, Ammonia concentration in portal and peripheral blood from CD- or LPD-fed SPF and GF mice, and from GF mice colonized with R. timonensis plus wild-type or ΔnrfA Bilophila. i, Ammonium chloride (NH4Cl) was administered in the drinking water to the indicated gnotobiotic mice, and iWAT Ucp1 and hepatic Fgf21 expression were assessed. j, Hepatocyte organoids derived from five human donors were stimulated with the indicated concentrations of NH4Cl, and FGF21 and CSAD mRNA expression was analysed by qPCR (n = 2 per group). Circles represent individual mice; data are mean ± s.d. Box plots show median, interquartile range and data range. Statistics: one-way ANOVA with Benjamini–Hochberg correction (b,c,e–j) and two-tailed Mann–Whitney test (d). NS, not significant.
To further assess the role of NrfA in WAT browning, we generated a nrfA-deficient Bilophila sp. 4_1_30 (St.14) mutant (Supplementary Fig. 7). This ∆nrfA mutant exhibited defective ammonia production in vitro, compared with the nrfA-sufficient (wild-type) Bilophila strain, which produced high levels of ammonia in protein-restricted settings (Extended Data Fig. 11b). In vivo, co-colonization of GF mice with the wild-type Bilophila strain and bile-acid-producing R. timonensis (St.31) induced significant browning, although the magnitude was substantially less than that observed with the complete hu4 strains (Fig. 6e). By contrast, co-colonization with R. timonensis and the ∆nrfA Bilophila mutant failed to induce browning (Fig. 6f and Extended Data Fig. 11d), despite comparable intestinal engraftment of the ∆nrfA and wild-type strains (Extended Data Fig. 11c). Notably, ∆nrfA Bilophila-colonized mice exhibited reduced hepatic expression of Fgf21 (Fig. 6g), whereas induction of hepatic Csad and Cyp39a1 expression and plasma bile acids was preserved or even enhanced (Extended Data Fig. 11e,f), suggesting that nrfA-mediated nitrogen metabolism specifically promotes Fgf21 induction, likely via ammonia production. Consistent with this, LPD feeding increased the levels of ammonia in the portal vein, but not in the peripheral blood, of SPF B6 mice and gnotobiotic mice colonized with wild-type Bilophila and R. timonensis. By contrast, no such increase was observed in GF mice or in mice colonized with the ∆nrfA strain (Fig. 6h). Supplementation with ammonium chloride rescued hepatic Fgf21 and iWAT Ucp1 and Elovl3 expression in mice colonized with R. timonensis and ∆nrfA Bilophila (Fig. 6i and Extended Data Fig. 11g). Moreover, stimulating human hepatocyte organoids with ammonia58 induced the expression of FGF21 in a dose-dependent manner without affecting bile acid biosynthesis genes (Fig. 6j). These results suggest that microbiota-derived ammonia enters the portal circulation and selectively induces hepatic FGF21 expression, thereby contributing to WAT browning.
Having elucidated the roles of human-derived, nitrogen-metabolizing Bilophila and bile-acid-modifying Romboutsia in browning, we revisited the mu20-mix. Two strains, Adlercreutzia sp. (St.1H8) and Parvibacter caecicola (St.1B6) were found to carry LSP-containing nrfA homologues and produced ammonia in vitro (Extended Data Fig. 12a,b). In addition, Faecalibaculum rodentium (St.1C4) was found to be capable of bile acid deconjugation as well as 7α- and 3α-dehydroxylation, producing CA, 7oxoDCA and 3oxoCA from tauro-CA (Extended Data Fig. 12c). We thus selected these three strains and combined them with Blautia sp. (St.27G3) and Turicibacter sp. (St.80E1). The resulting five-strain consortium (mu5-mix) robustly increased portal ammonia levels, hepatic Fgf21 mRNA expression and iWAT browning, comparable to the levels induced by the parental mu20-mix (Fig. 3k and Extended Data Fig. 12d). By contrast, exclusion of the two nrfA-positive strains (mu3-mix) or colonization with a random seven-strain mixture (mu7-mix) did not elicit these effects. These findings suggest that diverse microbial consortia, derived from either the human or the mouse microbiota, can promote adipose tissue browning in mice fed an LPD, provided that they possess both ammonia-producing and bile-acid-modifying capacities.
This study identifies low protein intake as a dietary context in which specific gut microorganisms engage coordinated host pathways to promote adipose tissue browning (see also Supplementary Discussion). By combining gnotobiotic models with defined human- and mouse-derived bacterial consortia, we show that distinct microbial communities can elicit a shared browning phenotype, provided that they encode complementary functional capacities of bile acid modification and ammonia production. These functions engage two host pathways: activation of FXR in adipose progenitor populations and induction of hepatic FGF21. These pathways probably act in parallel and are non-redundant, because disruption of either axis compromises browning, whereas activation of one alone is insufficient.
Our findings support a model in which protein scarcity is sensed by specific members of the microbiota, which respond accordingly, leading to increased levels of systemic bile acids and hepatic ammonia. This activates FXR and FGF21 signalling, which together drive sympathetic innervation, adipose remodelling and metabolic adaptation. Host–microbiota cooperation in nitrogen conservation and the maintenance of amino acid homeostasis through ammonia production represents an evolutionarily conserved adaptive strategy under conditions of protein scarcity59. This framework illustrates how the gut microbiota functions as an active interpreter of dietary composition, linking nutrient availability to coordinated host physiological responses.
Despite these advances, several key limitations remain. Mechanistically, it will be crucial to determine how the microbiota senses low-protein conditions; how the LPD-conditioned microbiota preferentially increases certain bile acids in the systemic circulation; how specific members of the microbiota influence multiple organs; whether and how FXR signalling promotes adipose progenitor differentiation; and how FGF21, FXR and possibly other factors lead to a substantial remodelling of sympathetic neurons. Although an LPD consistently induced metabolic changes, including weight loss and improved glucose tolerance, the directionality of the relationship between Ucp1 upregulation and weight loss is difficult to dissect. The specific contribution of beige adipocytes to overall metabolic effects needs to be addressed in future studies. Ucp1 mRNA expression is highly dynamic and does not necessarily indicate active thermogenesis. Indeed, reports suggest that the anti-obesity activity of FGF21 is mediated at least in part by UCP1-independent pathways60. As such, the LPD-activated pathways involving FGF21 induction, bile acid signalling, WAT browning and additional factors are likely to contribute to both UCP1-dependent and UCP1-independent metabolic outcomes, the quantitative deconvolution of which warrants further investigation.
SPF C57BL/6 (B6), BALB/c and ICR mice were purchased from Japan SLC, CLEA Japan and the Jackson Laboratory Japan. Male B6 mice aged 7 to 17 weeks were used unless otherwise specified. GF male B6 mice were purchased from Sankyo Labo Service Corporation and CLEA Japan. GF rederivation of SPF mutant mice was done at the gnotobiotic facilities of RIKEN and Keio University. In brief, in vitro fertilization (IVF) was used to generate embryos (typically using eggs and sperm from heterozygous pairs), which were then transplanted into IQI pseudo-pregnant female recipients. After embryo transfer, the recipient females underwent Caesarean sections on embryonic day 18. The intact uterine horns containing pups were passed through a germicidal bath, after which the pups were delivered into flexible plastic GF isolators and suckled by GF lactating foster mothers. This method enabled the generation of GF cohorts of 10–40 mice, all born on the same date (littermates or equivalents), thereby supporting consistent experimental conditions. Fgf21−/− mice were generated as previously described61. Triple-knockout mice lacking the β1-, β2- and β3-adrenergic receptors (Adrb1−/−Adrb2−/−Adrb3−/−) were used with permission from B. Lowell62. Il4−/− mice (G4 mice) were supplied by W. E. Paul63. R26:lacZbpAfloxDTA mice were provided by D. Riethmacher64 and subsequently crossed with Il5-cre or Lyz2-cree mice. Nr1h4−/−, Gpbar1−/−, Nr1h4fl/fl, Vil1-cre, Alb-cre, Adipoq-cre, Tcrb−/−, Tcrd−/−, Tbx21−/−, Rorc−/− (homozygous of Rorc(γt)-EGFP mice), Il5-cre and Lta−/− mice were purchased from the Jackson Laboratory. Rag2−/−Il2rg−/− mice were obtained from Taconic. Il33−/− mice were obtained from RIKEN BRC with permission from S. Nakae. Dpp4-cre (Dpp4-RFP, -cre) mice were obtained from RMRC. Unless otherwise indicated, mice were housed under controlled conditions, including a temperature range of 23–25 °C, a humidity of 40–60% and a 12-h light–dark cycle. In the experiment at thermoneutral conditions, SPF mice were kept in an incubator (MIR-154, PHCbi, Japan) at 30 °C according to previous work65. Magnetic resonance imaging (MRI) data collection was outsourced to PRIMETECH and collected using an M7 Compact MRI system (Aspect Imaging). Autoclaved water and gamma-irradiated (50 kGy) sterilized food were provided ad libitum throughout the experiments. Unlike autoclaving, gamma irradiation avoids heat-induced alterations in food components. The specific diets and durations of each experiment are detailed in the corresponding figure panels. Mice were randomly allocated into experimental groups. All animal experiments were approved by the Institutional Animal Care and Use Committee of Keio University and the RIKEN Yokohama Institute.
All experimental diets were obtained from Research Diets or Oriental Yeast. The control diet used in this study was a 50-kGy-irradiated AIN-93G diet (product D19090404) containing 20 kcal% protein (mineral acid casein), 64 kcal% carbohydrate and 16 kcal% fat. Unless otherwise specified, the LPD was an isocaloric AIN-93G-based diet containing 7 kcal% protein, 77 kcal% carbohydrate and 16 kcal% fat (product D20121501; see Supplementary Tables 3 and 4). Diets with varying proportions of protein, carbohydrate and fat were formulated based on the AIN-93G diet, with the exception of the ketogenic diet and the ketogenic–control diet, which have a lower fat content than AIN-93G does. The ketogenic diet and ketogenic–control diets (products D20012303 and D20012304 from Research Diets, respectively) use cocoa butter as the main fat source, with the protein concentration adjusted to 20 kcal% to match the AIN-93G diet. Because the ketogenic–control diet contains only 10% fat, which is lower than the fat content of AIN-93G, it is referred to as the low-fat diet in Fig. 1a. For experiments using defined amino acid diets, natural protein (mineral acid casein) was replaced with pure amino acids, aligned with the amino acid composition of the protein. The total amino acid concentration was then adjusted, ranging from 20 to 2.5 kcal%, or set to 2.5% kcal content for individual EAAs, while maintaining the levels of other amino acids at 20%, and isocaloric conditions were maintained by varying the amount of carbohydrate (see Supplementary Table 5).
SPF C57BL/6 mice were at first fed an HFD for 2, 4 or 9 weeks. For HFD-to-CD versus HFD-to-LPD experiments, HFD32 (CLEA Japan; 60 kcal% fat and 20 kcal% protein) was used. For the HFD-to-HF/LPD versus HFD-to-HFD experiments, an AIN-93G-based HFD (32 kcal% fat and 20 kcal% protein) was used (Supplementary Table 6). Mice were then switched to one of the following diets for 4 or 6 weeks: control diet (20% fat and 20% protein), LPD (20 kcal% fat and 7 kcal% protein) or HF/LPD (32 kcal% fat and 7 kcal% protein). Quantification of plasma ALT, cholesterol and triglycerides was outsourced to Oriental Yeast. Oral glucose tolerance tests (OGTTs) were performed 7 days before euthanasia. For the hu4-mix treatment experiments, GF mice were inoculated with faecal microbiota from human participant T07 and fed a HFD (HFD32) for four weeks, then switched to an LPD. On days 0 and 3 after the diet switch, mice were given two oral doses of the hu4 strains or vehicle control. After 6 weeks on the LPD, mRNA expression of Ucp1 in iWAT and BAT, iWAT mass, hepatic Fgf21 mRNA expression and plasma levels of cholesterol, triglycerides and ALT were assessed. An OGTT was performed after 3 weeks of LPD feeding.
Faecal caloric content was measured using bomb calorimetry, as previously described66. In brief, faecal samples were dried overnight at 60 °C, weighed and analysed for energy content using a bomb calorimeter (Parr 6100EA with Semimicro Bomb) calibrated with standard benzoic acid (6,320 cal g−1).
Cold exposure and treatments with a β3-adrenergic receptor agonist, tungsten, ammonium chloride and recombinant FGF21
For cold exposure, mice were housed at 6 °C for 7 days. For treatment with a β3-adrenergic receptor agonist, mice were administered CL316,243 daily by intraperitoneal injection (20 μg per mouse per dose, SIGMA C5976) for seven consecutive days. For tungsten treatment, mice were provided with 0.22-μm filter-sterilized sodium tungstate dihydrate (Na2WO4/2H2O, Nacalai Tesque 32011-25) in their drinking water at concentrations of 0.1% or 0.5% for 2 weeks. Throughout the treatment period, mice had ad libitum access to the tungsten-containing water, and both the remaining water volume and the health status of the mice were carefully monitored. On the basis of daily observations, mice in the 0.1% tungsten treatment group did not exhibit any noticeable changes in food or water consumption. By contrast, those in the 0.5% tungsten group showed signs of adverse effects, including mildly reduced appetite. For ammonium chloride (NH4Cl) treatment, mice were given 0.22-μm filter-sterilized NH4Cl (Nacalai Tesque 02423-65) in their drinking water at concentrations of 5 or 50 µg ml−1 for 5 weeks, corresponding to the full duration of LPD feeding. Administration of NH4Cl at either concentration did not result in any noticeable changes in food or water intake. For recombinant FGF21 treatment, GF B6 mice were fed a control diet for 1 week. On days 5, 6 and 7, they received intraperitoneal injections of recombinant human FGF21 (12 μg per mouse, twice daily, BioLegend 553804) or PBS.