- Yancheng E. Li (李妍成) ORCID: orcid.org/0000-0002-7110-24481,
- S. Francesca Antillon2,3,
- Grace F. Baron ORCID: orcid.org/0009-0008-3396-31171,
- Karthik Chamakura ORCID: orcid.org/0000-0002-6196-85712,3,
- Ry Young2,3 &
- William M. Clemons Jr ORCID: orcid.org/0000-0002-0021-889X1,4
Nature (2026)Cite this article
- 48 Altmetric
- Metrics details
- Antibiotics
- Cryoelectron microscopy
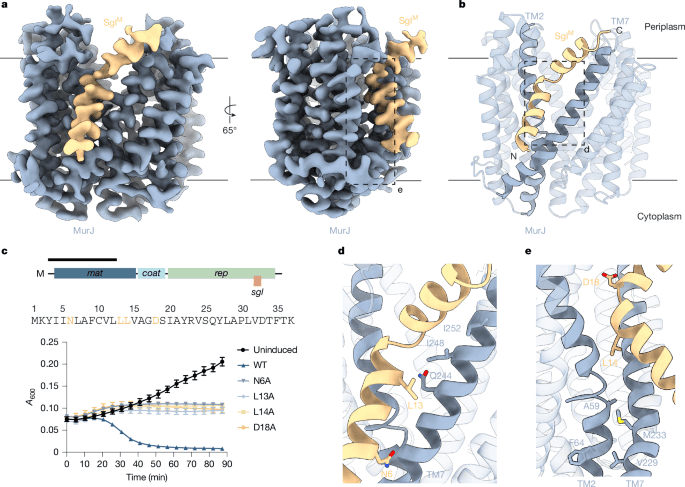
Antimicrobial drug resistance poses a global health challenge that necessitates the identification of new druggable targets1,2,3. The essential lipid II flippase MurJ is a promising yet underexplored antimicrobial target in bacterial cell wall biosynthesis4,5,6,7. The only known inhibitors of Gram-negative (diderm) MurJ are the single-gene lysis proteins (Sgls) from the lytic single-strand RNA phages M (SglM) and PP7 (SglPP7)8,9. SglM and SglPP7 have distinct evolutionary origins and share no sequence similarity. Here we describe a common mechanism of MurJ inhibition by these phage-encoded Sgls. We determined the structures of MurJ-bound SglM and SglPP7 and discovered a third distinct MurJ-targeting Sgl from the predicted phage Changjiang3 (SglCJ3) that we also characterized structurally. Our findings demonstrate that all three Sgls evolved convergently to trap MurJ in a periplasm-open conformation through a common MurJ interface, revealing a pathway for drug design.
This is a preview of subscription content, access via your institution
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
Receive 51 print issues and online access
- Purchase on SpringerLink
- Instant access to the full article PDF.
Prices may be subject to local taxes which are calculated during checkout
- Learn about institutional subscriptions
- Read our FAQs
- Contact customer support