DICER is an evolutionarily conserved RNase III enzyme that has a central role in RNA silencing, a crucial gene regulatory mechanism that influences diverse biological processes. DICER processes precursor microRNAs (pre-miRNAs) and long double-stranded RNAs (dsRNAs) into small regulatory RNAs, microRNAs (miRNAs) and small interfering RNAs (siRNAs), which target mRNAs for degradation or translational repression through the RNA-induced silencing complex (RISC)1,2,3. This cleavage activity also drives short-hairpin RNA (shRNA) technology, in which synthetic shRNAs are processed by DICER into siRNA duplexes that mimic endogenous RNAi pathways5,6,7,8,9,10.
Over the past two decades, studies have elucidated key aspects of DICER’s cleavage mechanism, including 5′- and 3′-end counting rules that determine cleavage sites around 21–22 nucleotides (nt) from the RNA ends4,11,12,13,14,15,16,17. Structural studies have shown that the 3′-end binds within a conserved pocket in DICER’s PAZ domain, whereas the 5′-end interacts with a pocket in DICER’s platform domain4,14,17,18,19,20,21,22,23,24,25. In human, mouse and fly DICER enzymes, the 3′-end terminal phosphate interacts with a cluster of conserved tyrosines and basic residues4,17,18,19,21,22. Similarly, plant DICER-like enzymes (for example, Arabidopsis thaliana DCL1 and DCL3) have a conserved 3′-end docking pocket, highlighted by aligned conserved tyrosines, basic residues, and phenylalanine24,25. These findings suggest that the 3′-end binding pocket is conserved across DICER and DICER-like enzymes.
The previous effort to identify the 5′-end binding pocket in DICER used crystal structures of its PAZ–platform–connector helix cassette bound to siRNAs, revealing a phosphate-binding pocket17. However, these structures might not represent the dicing state. A cryo-electron microscopy (cryo-EM) study resolved DICER in the dicing state, detailing a 5′-end binding pocket for 5′-U, involving R790, R821 and R10034. This pocket disfavours guanosine (5′-G), owing to steric clashes with the arginine residue (R821), suggesting that 5′-G reduces cleavage accuracy4. Similarly, a dicing-state structure of fly Dcr-1 identified a 5′-end binding pocket for 5′-U, with R1027 (conserved with R821 in human DICER) located in this pocket21. In A. thaliana DCL3, the 5′-end binding pocket consists of histidine and arginine residues, which are not conserved with human DICER25. By contrast, the 5′-end binding pockets of mouse DICER, fly Dcr-2 and Arabidopsis thaliana DCL1 remain undefined19,22,23,24. These findings suggest that, unlike the conserved 3′-end binding pocket, the 5′-end binding pocket varies among enzymes and is challenging to determine. Moreover, structural studies of human DICER and fly Dcr-1 have relied on RNA substrates with fixed 5′-U, and the effects of diverse RNA ends have not been examined.
DICER uses fine-tuning mechanisms, such as sequence motifs (for example, mWCU, YCR and GYM), bulges, loops and mismatches, to achieve precise cleavage at either position 21 (DC21) or position 22 (DC22) from the 5′-end6,9,26,27,28,29,30,31,32,33,34,35,36,37. However, the interaction between these features and end-binding rules, as well as how DICER coordinates RNA motifs with end-binding preferences for precise cleavage, remain unclear.
In this study, using massively parallel dicing assays and single-particle cryo-EM, we show that 5′-G, contrary to previous reports, enhances cleavage precision at DC21 for many substrates. Cryo-EM reconstructions uncover a previously unrecognized G-favoured binding pocket, distinct from the U-favoured binding pocket, directing cleavage to DC22. This dual-pocket mechanism reshapes our understanding of small-RNA biogenesis, showing how DICER integrates end-binding rules with RNA features for precise cleavage. RNA motifs such as mWCU and YCR cooperate with the 5′-end binding rule to refine specificity, and conflicts induce RNA conformational changes that override end-binding preferences. Cryo-EM further reveals dynamic rearrangements of the dsRBD and PAZ domains, repositioning RNA for precise catalysis and providing a framework for RNA processing in silencing pathways.
DICER cleavage accuracy is enhanced by 5′-G
When testing pre-mir-517a, a pre-miRNA with a 5′-C (pre-mir-517a_CU), we observed cleavage at both the DC21 and the DC22 site (Fig. 1a–c). To assess the contribution of the 5′-nucleotide (5′-nt), we replaced the 5′-C with other nucleotides and examined dicing outcomes. Substituting 5′-C with 5′-A (pre-mir-517a_AU) or 5′-G (pre-mir-517a_GU) shifted cleavage predominantly to DC21, with 5′-G showing the highest accuracy, whereas 5′-U (pre-mir-517a_UU), like 5′-C, supported cleavage at both sites (Fig. 1b,c and Extended Data Fig. 1a). Because 5′-G and 5′-A pair with U on the 3′-strand, whereas 5′-C and 5′-U do not, these substitutions altered the overhang geometry. To isolate the effect of the 5′-nt from the overhang context, we tested pre-mir-517a variants with comparable overhangs. Notably, G–U (pre-mir-517a_GU) showed higher DC21 accuracy than U–G, whereas A–U favoured DC21 and U–A favoured DC22 (Fig. 1b,c). These findings refute the hypothesis that 5′-G reduces cleavage accuracy4, and reveal that the 5′-nt strongly influences DICER’s cleavage-site preference: 5′-G and 5′-A promote DC21, whereas 5′-C and 5′-U favour DC22.
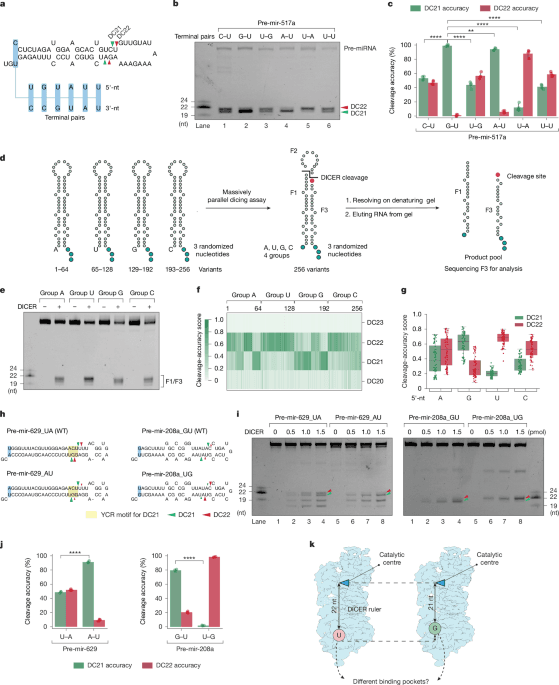
a,Schematic of pre-mir-517a substrates, illustrating terminal pairs (for example, G–U and C–U) at the 5′- and 3′-ends. Green and red arrowheads indicate DC21 and DC22 cleavage sites, located 21 and 22 nt from the 5′-end, producing fragments F1, F2 and F3. b, In vitro dicing assays for pre-mir-517a and variants using 2 pmol RNA and 1 pmol DICER. F1 and F3 fragments are 22 nt (DC22) or 21 nt (DC21), whereas F2 is undetectable on gels. c, Cleavage accuracy of DICER at DC21 and DC22 for pre-mir-517a variants, based on four independent experiments. Statistical significance was determined using a two-tailed, two-sample t-test (**P < 0.01, ****P < 0.0001). d, Schematic of massively parallel dicing assays using pre-mir-324 groups with fixed 5′-nt (A, G, U or C) and randomized 3′-overhang. Cleaved products (F3) were cloned, sequenced and analysed. e, Cleavage patterns from pre-mir-324 groups with distinct 5′-end nucleotides. f, Cleavage sites (DC20–DC23) identified from sequencing results. n = 256 variants. g, Cleavage accuracy at DC21 and DC22 for pre-mir-324 groups, calculated from sequencing data. n = 64 for each comparing group. h, Diagrams of pre-mir-629, pre-mir-208a and variants, showing terminal pairs. Green and red arrowheads indicate DC21 and DC22 cleavage sites. i, In vitro dicing assays for pre-mir-629, pre-mir-208a and variants using 2 pmol RNA and 1 pmol DICER. F1 and F3 fragments are 22 nt (DC22) or 21 nt (DC21); F2 is undetectable. Green arrowheads, DC21; red arrowheads, DC22. j, Cleavage accuracy of DICER at DC21 and DC22 for pre-mir-629 and pre-mir-208a variants, based on three independent experiments. Statistics as in c. k, Mechanistic model illustrating how 5′-nt affects DICER cleavage. Guanine (G) and adenine (A) favour DC21 cleavage, whereas uracil (U) and cytosine (C)—especially U—favour DC22 cleavage.
Parallel dicing assays confirm the effects of the 5′-nt
To generalize these findings, we performed massively parallel dicing assays using pre-mir-324, cleaved by DICER at DC20, DC21 and DC2238,39,40. We synthesized four groups of pre-mir-324 variants, each with a specific 5′-nt and randomized 3′-ends, producing 64 unique sequences per group (Fig. 1d). After DICER cleavage, F3 fragments were sequenced to identify cleavage sites (Fig. 1e and Extended Data Fig. 1b). The assays were highly reproducible, with strong correlation between replicates (Extended Data Fig. 1c,d). Sequencing confirmed that DICER cleaves pre-mir-324 mainly at DC21 and DC22 (Fig. 1f). Whereas the 3′-nt had no detectable effect in our randomized pre-mir-324 context (Extended Data Fig. 1e), the 5′-nt strongly influenced site preference: 5′-G yielded the highest DC21 accuracy, followed by 5′-A, whereas 5′-C and 5′-U favoured DC22 (Fig. 1g). These results confirm that 5′-G enhances, rather than impairs, cleavage accuracy by promoting DC21 cleavage, underscoring the prominent role of the 5′-nt in determining DICER’s cleavage-site preference.
Effects of the 5′-nt validated across pre-miRNAs
To validate the role of the 5′-nt in cleavage specificity, we synthesized pre-mir-324 variants with a fixed 3′-end and varying 5′-nts (Extended Data Fig. 1f). Consistent with the massively parallel assays, 5′-G and 5′-U promote cleavage at DC21 and DC22, respectively, whereas 5′-A and 5′-C support cleavage at both sites, with slight preferences for DC21 and DC22, respectively (Extended Data Fig. 1g,h). To isolate the influence of the 5′-nt, we generated variants with similar 2-nt overhang structures but differing base pairs (A–U versus U–A, or G–U versus U–G), confirming that 5′-G strongly favours DC21, 5′-U favours DC22 and 5′-A promotes DC21 more than 5′-U but less than 5′-G (Extended Data Fig. 1i–k). Finally, variants with similar 3′-nt overhangs but differing 5′-nts showed that 5′-A favoured DC21, whereas 5′-U and 5′-C supported DC22, with 5′-U showing the strongest bias (Extended Data Fig. 1l–n).
To test whether the effects of the 5′-nt apply to other pre-miRNAs, we examined pre-mir-629 and pre-mir-208a, which naturally have 5′-U and 5′-G, respectively (Fig. 1h). In pre-mir-629, DICER cleaves at both DC21 and DC22, with DC21 cleavage driven by a YCR motif. Changing 5′-U to 5′-A reduces DC22 cleavage, confirming that 5′-U enhances DC22 cleavage (Fig. 1i,j). Similarly, pre-mir-208a shows DC21 cleavage with 5′-G and DC22 cleavage with 5′-U (Fig. 1i,j). These findings show that 5′-G promotes DC21, whereas 5′-U promotes DC22 (Fig. 1k).
Cryo-EM shows DICER bound to 5′-G and 5′-U shRNAs
To examine how 5′-G and 5′-U influence DICER cleavage specificity at DC21 and DC22, we solved cryo-EM structures of DICER bound to 5′-G (26S-GU) or 5′-U (26S-UG) shRNAs (Fig. 2a,b). Adding Ca2+ during reconstitution trapped the dicing state by inhibiting catalysis while preserving the cleavage-ready conformation. Cryo-EM maps of the DICER–26S-GU and DICER–26S-UG complexes were obtained at 3.34-Å and 3.37-Å resolution, respectively, as determined by gold-standard Fourier shell correlation (GS-FSC) (Extended Data Fig. 2a–f and Supplementary Table 1). Multiple rounds of two-dimensional (2D) classification selected particles that resembled the dicing-state complex, consistent with previous studies4 (Extended Data Fig. 2a,b). A bias in particle orientation was observed, which is likely to be due to DICER’s asymmetric shape and mass distribution, which might reduce directional resolution and lead to anisotropic reconstructions (Extended Data Fig. 2f). Local-resolution analysis showed slightly higher resolution in protein densities than in RNA densities (Extended Data Fig. 2d). Using the published structural model4 (Protein Data Bank (PDB): 7XW2), we refined atomic structures of DICER in complex with both shRNAs, revealing detailed architecture and domain organization (Extended Data Fig. 2g). The 26S-UG and 26S-GU RNAs included mWCU and YCR motifs, promoting cleavage at DC2235,36, which allowed us to investigate the effects of 5′-nt and RNA motifs on DICER–RNA interactions (Fig. 2c and Extended Data Fig. 3a).
a, Schematic of human DICER domains: DUF283, PAZ, RIIIDa and RIIIDb (RNase III) and dsRBD, with amino acid boundaries labelled. b, Sequences of the 26S-UG and 26S-GU shRNAs that were used in cryo-EM studies. The mWCU and YCR motifs guide DC22 cleavage. c, Cleavage accuracy of DICER for 26S-UG and 26S-GU calculated from three independent assays, showing precise cleavage at DC22. d, Cryo-EM reconstructions of DICER–26S-UG and DICER–26S-GU complexes, with domains colour-coded. e, Structural comparison of DICER complexes reveals conformational shifts in dsRBD and PAZ after RNA binding. RMSD values show significant movement compared with apo-DICER (PDB: 7XW3) and DICER–pre-let-7a-1GYM (PDB: 7XW2). f, Alignment of cryo-EM densities shows that shRNA-bound complexes are more compact (57.7–58.5 Å) than DICER/pre-let-7a-1GYM (68.0 Å). g, PAZ and dsRBD translocate in the DICER–26S-UG and DICER–26S-GU complexes relative to DICER–pre-let-7a-1GYM, with RMSD values of 7.4 Å (PAZ) and around 5.0 Å (dsRBD), aligning the RNA substrate with the catalytic centres. h, PAZ adopts an ‘inner’ conformation in DICER–shRNA complexes, bending the 3′-overhang, unlike the ‘outer’ conformation in DICER–pre-let-7a-1GYM. i, Buried surface area analysis shows increased RNA–DICER interaction in shRNA-bound states, compared with DICER–pre-let-7a-1GYM. j, Proposed model: the PAZ-dsRBD ‘in mode’ aligns the RNA backbone for precise cleavage, whereas the ‘out mode’ can cause RNA misalignment.
The cryo-EM structures reveal the organization of DICER’s key domains, including the platform, PAZ, RIIIDa, RIIIDb, dsRBD, connector helix and bound RNA (Fig. 2d). Partial densities for helicase domains, probably Hel1, are also observed (Extended Data Fig. 3b). However, as in previous studies4,22, incomplete densities are seen for helicase residues (1–564), DUF283 (590–715) and certain loops in RIIIDa and RIIIDb (1389–1545 and 1588–1658). In both the DICER–26S-GU and the DICER–26S-UG complex, the shRNAs are fully docked at the catalytic centre, formed by intramolecular dimerization of RIIIDa and RIIIDb. The alignment of conserved acidic residues in RIIIDa (E1316, D1320, D1561 and E1564) and RIIIDb (E1705, D1709, D1810 and E1813) with the RNA cleavage site, along with calcium ions positioned between these residues and the RNA, confirms the dicing-ready conformation (Extended Data Fig. 3c).
dsRBD and PAZ domain conformational changes
To examine how DICER adapts during RNA binding and cleavage, we compared our RNA-bound dicing-state structures (DICER–26S-GU and DICER–26S-UG) with the previously reported dicing-state structure (PDB: 7XW2) and apo-DICER (PDB: 7XW3)4. Root-mean-square deviation (RMSD) analysis, measuring atomic displacement between corresponding residues, revealed significant structural variability in the dsRBD and PAZ domains, highlighting transitions between functional states (Fig. 2e). Although the overall conformational changes in the dsRBD are consistent with previous observations4 (7XW2), the inward motion of the PAZ domain is distinctly observed in our RNA-bound dicing-state structures.
Alignment of the RNA-bound structures (DICER–26S-GU and DICER–26S-UG) with the 7XW2 dicing-state structure reveals a more compact conformation in the RNA-bound states, with a narrower overall width (57.7–58.8 Å for DICER–26S-GU and DICER–26S-UG vs. 68.0 Å for 7XW2) (Fig. 2f). This compaction is driven by structural rearrangements in the PAZ domain, which shifts inwards by around 7.4 Å after RNA binding (Fig. 2g). Specifically, the α-helix (residues 968–976) that anchors the RNA 3′-end moves inward by 7.6–8.1 Å, and adjacent β-sheets shift by 5.0–5.3 Å (Extended Data Fig. 3d). This differential motion compresses the PAZ domain, and probably contributes to bending nucleotides near the 3′-end (Fig. 2h).
The dsRBD and PAZ domain move in coordination
Structural comparisons of our RNA-bound dicing-state structures (DICER–26S-UG and DICER–26S-GU) with the previously reported dicing-state structure (7XW2) reveal coordinated movements of the dsRBD and PAZ domains. Although the dsRBD undergoes similar changes in all RNA-bound states, compared with apo-DICER (Fig. 2e), it shifts around 5 Å closer to DICER’s longitudinal axis in the DICER–26S-UG and DICER–26S-GU structures compared with 7XW2 (Fig. 2g). Combined with the inward motion of the PAZ domain, this shift increases the buried surface area of the DICER–RNA complex (Fig. 2i). These rearrangements align the RNA duplex along DICER’s axis, optimizing the positioning of cleavage sites for efficient RNA processing (Fig. 2i).
We propose that the dsRBD and PAZ domains act like chopsticks, gripping and aligning RNA substrates for precise double cleavages on the 5′- and 3′-strands (Fig. 2j). In our shRNA-bound structures, the RNA duplex remains tightly aligned without expansion (Extended Data Fig. 3e). By contrast, in pre-let-7a-1GYM from 7XW2, the dsRBD and PAZ domains are farther from the RNA duplex, which would require RNA helix widening to align both cleavage sites with the catalytic centre for accurate cleavage (Extended Data Fig. 3f).
Conserved structure of 3′-end binding pockets
In our dicing-state structures, the terminal wobble pairs (G–U and U–G) are unpaired, allowing the 3′-nt and 5′-nt to interact with the PAZ and platform domains, respectively (Fig. 3a,b, surface-view panels). The 3′-nt consistently occupies the conserved 3′-end binding pocket, in which the terminal phosphodiester linkage interacts with conserved residues, including tyrosines (Y936, Y971, Y972 and Y976) and basic residues (R937 and K975) (Fig. 3a,b, 3′-end binding pocket panels). This configuration aligns with the results of previous structural studies4,14,19,21,22,23,24, confirming the conserved nature of the 3′-end recognition mechanism of DICER. Notably, no residues interact with the terminal base of the 3′-nt, indicating that its identity is unlikely to affect cleavage-site selection (Extended Data Fig. 1e).
a, Binding pockets in the DICER–26S-UG complex. Left, cryo-EM model of DICER bound to 26S-UG with colour-coded domains. Left middle, electrostatic surface view of the PAZ–platform region, showing the paths of the 5′- and 3′-ends. Right middle, close-up view of the 3′-end pocket, with A64 and labelled residues. Right, 5′-end pocket, showing U1 contacts with R821 and R1003. b, Binding pockets in the DICER–26S-GU complex. Panels as in a. The 3′-end pocket accommodates A64, whereas the 5′-end pocket binds to G1, with residues D991 and H992 indicated. c, The alignment of the RNA backbone is influenced by 5′-end binding. In DICER–26S-GU, the RNA backbone shifts one base pair upwards towards the catalytic centre, compared with DICER–26S-UG. d,e, In vitro dicing of pre-mir-517a_GU (d) and pre-mir-517a_CU (e) by wild-type (WT) and mutant DICER. Left, denaturing gels showing DC21 and DC22 products. Middle, quantification of cleavage accuracy; significance by two-tailed, two-sample t-test (***P < 0.001, ****P < 0.0001; NS, not significant). Right, pre-miRNA schematics with cleavage sites. f, Cryo-EM map of DICER(D991G/H992G)–26S-GU, with colour-coded domains and RNA duplex (26S-GU) indicated. g, Electrostatic surface view of the active-site region of the mutant DICER, highlighting the bound duplex and the 5′- and 3′-strand positions. h, Close-up view of the PAZ domain and the terminal pair G1–U62. i, Proposed mechanism: distinct 5′ binding pockets (5′-G-favoured for DC21 and 5′-U-favoured for DC22) determine cleavage-site selection.
Identification of a 5′-G-favoured binding pocket
Unlike the conserved 3′-end binding pocket, the 5′-nt shows distinct binding differences between the DICER–26S-UG and DICER–26S-GU structures (Fig. 3a,b, surface-view panels). In the DICER–26S-UG structure, the 5′-U occupies the previously identified pocket4, in which R1003 interacts with the 5′-phosphate and R821 recognizes the uridine base (Fig. 3a, 5′-end binding pocket panels, and Extended Data Fig. 4a). However, R790, which was previously thought to interact with uridine, shows no interaction, consistent with mutation studies suggesting that R790 is not essential for uridine recognition4.
Our DICER–26S-GU structure reveals a previously unidentified 5′-G binding pocket. Here, the 5′-phosphate of 5′-G is anchored by H992, while its guanine base forms hydrogen bonds with D991 (Fig. 3b, 5′-end binding pocket panels, and Extended Data Fig. 4a). D991 and H992 create a G-selective microenvironment that is distinct from the 5′-U site. Binding of 5′-G shifts the RNA by approximately one nucleotide upwards along the hairpin’s lower stem compared with 5′-U (Fig. 3c). With identical upper RNA sequences, this shift positions DC21 at DICER’s catalytic centre for 5′-G, whereas 5′-U positions DC22. These structural differences provide a mechanistic basis for how 5′-nt identity biases cleavage-site selection.
Validation of the 5′-G binding pocket
To confirm the functional importance of the 5′-G binding pocket, we mutated the key residues D991 and H992 (Extended Data Fig. 4b). The mutations D991G, H992G and D991G/H992G reduced DC21 cleavage in pre-miRNAs with 5′-G, showing that these residues are essential for the function of the 5′-G binding pocket (Fig. 3d,e and Extended Data Fig. 4c–e). These mutations did not affect DC22 cleavage in pre-miRNAs with 5′-U, confirming the pocket’s specificity for 5′-G substrates (Fig. 3d,e and Extended Data Fig. 4c–e). Although the overall cleavage efficiency remained unchanged, the mutations influenced site selection, suggesting that these residues have a role in adjusting DICER positioning between DC21 and DC22. We further expressed pre-mir-517a_GU or pre-mir-517a_UG in HCT116 DICER-knockout cells with wild-type DICER or DICER(D991G/H992G), and performed miRNA sequencing (Extended Data Fig. 4f). Wild-type DICER produced more cleavage at DC21 from pre-mir-517a_GU than from pre-mir-517a_UG, consistent with the in vitro findings. However, the D991G/H992G mutant yielded less DC21 cleavage from pre-mir-517a_GU than wild-type DICER, but similar levels of DC21 and DC22 cleavage from both variants, confirming that DC21 enhancement by 5′-G depends on D991 and H992 (Extended Data Fig. 4g). In cells, cleavage at DC22 predominates, in part owing to the influence of the cofactor TRBP (transactivation response element RNA-binding protein) (Extended Data Fig. 4h).
We determined the structure of the DICER(D991G/H992G) mutant bound to 26S-GU (Fig. 3f and Extended Data Fig. 5a–g). The map shows continuous density for the loop containing residues 991–992 but lacks side-chain density for D991 or H992, consistent with their substitution by glycine (Extended Data Fig. 5h). Unlike in the wild-type DICER–26S-GU complex, the 5′-G shifts from the 5′-G pocket towards the 5′-U pocket, but does not interact with R1003 or R821 as 5′-U does (Fig. 3g and Extended Data Fig. 5i). Instead, the 5′-G is retained by base pairing with the opposing U (Fig. 3h and Extended Data Fig. 5j). The 3′-nt remains in the conserved 3′-end pocket (Extended Data Fig. 5k). These findings provide further evidence for the role of D991 and H992 in forming the 5′-G binding pocket.
The 5′-G binding pocket is conserved across species
Sequence analysis shows that D991 and H992, forming the core of the 5′-G binding pocket, are conserved across DICER enzymes in several species (Extended Data Fig. 6a), suggesting that this is an evolutionarily conserved functional feature. Massively parallel dicing assays on fly Dcr-1, a homologue of human DICER, revealed similar cleavage specificity, with 5′-G promoting DC21 cleavage and 5′-U favouring DC22 (Extended Data Fig. 6b–f). Consistently, an analysis of miRNAs derived from pre-miRNAs with different 5′-nts (A, G, U or C) across species showed that 5′-G pre-miRNAs yield the most DC21 miRNAs, whereas 5′-U pre-miRNAs produce the most DC22 miRNAs (Extended Data Fig. 6g,h).
On the basis of these findings, we propose a dual-pocket model for 5′-end recognition in DICER: a G-favoured binding pocket (formed by D991 and H992) for DC21 cleavage, and a U-favoured binding pocket (stabilized by R1003 and R821) for DC22 cleavage. These pockets align the RNA substrate 21 or 22 nucleotides from the 5′-end with DICER’s catalytic centres, enabling flexible yet precise RNA processing (Fig. 3i).
Resolving 5′-end counting and motif rules
In the DICER–26S-UG structure, the 5′-U directs cleavage at DC22, consistent with mWCU–YCR motifs that also guide DC22 cleavage. By contrast, the DICER–26S-GU structure shows that the 5′-G directs cleavage at DC21, conflicting with the DC22-guiding mWCU–YCR motifs. However, despite this conflict, 26S-GU is ultimately cleaved at DC22 (Fig. 1c). To understand how DICER resolves this conflict, we compared the protein structure of DICER in the DICER–26S-GU and DICER–26S-UG structures. The overall protein architectures are nearly identical, with only minor adjustments depending on whether the 5′-nt is G or U (Fig. 4a). Specific changes occur in a helix near the 3′ cleavage site in RIIIDa, the PAZ helix and a loop in the 5′-end binding pocket (Extended Data Fig. 7a). These subtle shifts suggest that DICER resolves cleavage conflicts mainly through local rearrangements, rather than through large-scale conformational changes.
a, Structural comparison of the DICER–26S-UG and DICER–26S-GU complexes. RMSD heat maps show minor structural differences in the 5′ binding pocket, PAZ domain and RIIIDa near the 3′ cleavage site. b, Alignment of the 26S-UG and 26S-GU RNA structures with their corresponding structures from AlphaFold3 (AF3)-predicted models. c, RNA flexibility and DICER’s dsRBD interactions: the 26S-GU complex exhibits distortion of the RNA backbone, aligning DC22 with the catalytic centre. d, Pre-dicing state of DICER bound to pre-mir-517a_GU. Cryo-EM density shows the PAZ domain (gold) engaging the duplex end and the dsRBD (gold) positioned along the RNA duplex (green), with other domains in grey. e, Dicing state. The duplex (green) docks into RIIIDa and RIIIDb (RIIIDa/b) as the PAZ domain and dsRBD (red) reposition to clamp the RNA for cleavage. f, Model of domain motion. Arrows indicate the movements of the dsRBD and the PAZ domain during activation. g, Density view of the end of the RNA duplex, showing G1 and opposing U59. h, The RNA duplex 5′-end occupies the DC21 pocket; the boundary loop (red) lies between DC21 and DC22. Open circles indicate pocket positions. i, Close-up view of the 5′-end region, showing G1 and the 5′-phosphate (PO4) coordinated near residues D991 and H992. j, RNA helical geometry comparison, highlighting the duplex distortions in 26S-GU compared with AlphaFold3 and experimental models of pre-mir-517a_GU. k, Proposed mechanism. 5′-U anchor: the 5′-U pocket aligns the mWCU–YCR motif at DC22 for cleavage. 5′-G anchor: the 5′-G pocket favours DC21 cleavage, misaligned with the mWCU–YCR motif at DC22. Thus, motif-driven RNA conformational changes override the 5′-G rule, distort the RNA backbone and ensure DC22 cleavage.
Our analysis reveals that RNA undergoes substantial conformational changes upon interaction with DICER, and these are particularly evident in the DICER–26S-GU structure compared with DICER–26S-UG. Although the predicted secondary structures of 26S-GU and 26S-UG are similar (AlphaFold341; Fig. 4b), DICER induces distinct structural changes at the RNA ends and cleavage sites (Fig. 4b). At the ends, interactions with the 5′- and 3′-end binding pockets destabilize the base pairs of terminal nucleotides, anchoring them in specific positions4,14 (Fig. 4b). Near the cleavage site, interactions between RNA motifs (for example, mWCU and YCR) and DICER shape structural distortions. Although YCR does not directly interact with DICER residues, R1855 in dsRBD engages with the 19-CC mismatch of mWCU, aiding RNA backbone adjustments for cleavage positioning (Fig. 4c).
In the DICER–26S-UG structure, the 5′-U pocket positions DC22 at the catalytic centre, with a moderate separation between the 19-CC mismatch and R1855. In the DICER–26S-GU structure, the 5′-G pocket shifts the backbone upwards, at first aligning DC21 and bringing the 19-CC mismatch closer to R1855. Engagement of this motif imposes a register-specific strain: the 26S-UG backbone undergoes a subtle adjustment, whereas the 26S-GU backbone distorts more substantially to accommodate the interaction and realign DC22 at the catalytic centre (Fig. 4c and Extended Data Fig. 7b,c). This mechanism might help to explain how RNA-motif interactions can override the DC21 bias of the 5′-G pocket, favouring DC22.