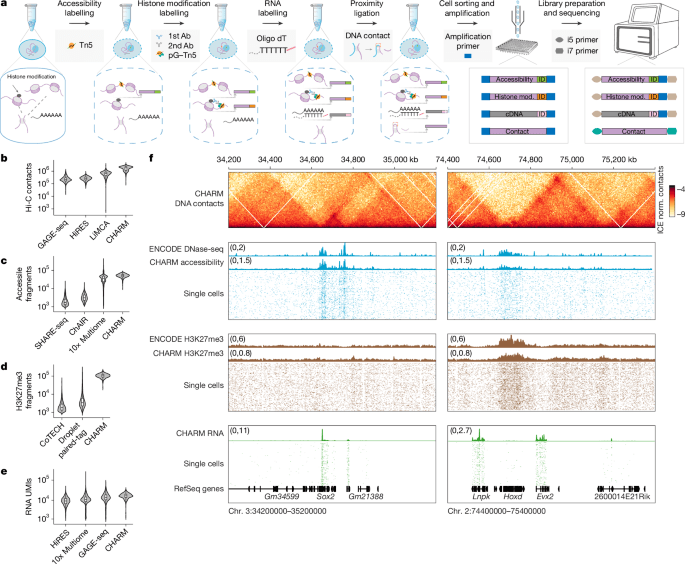
In a landmark advance for genomics, an international team of researchers has developed the first method to simultaneously profile four critical layers of gene regulation—genome conformation, histone modifications, chromatin accessibility, and gene expression—at single-cell resolution. Published in Nature, this four-omics technique, called CHARM (Chromatin Accessibility and Regulatory Multi-omics), enables scientists to dissect how DNA folding and epigenetic marks coordinate to control which genes are turned on or off in individual cells. The breakthrough offers new tools to understand development, disease, and aging by revealing the dynamic interplay between genome structure and function that shapes cellular identity.
- The study introduces CHARM, a single-cell four-omics sequencing method combining genome conformation, histone modifications, chromatin accessibility, and gene expression.
- Researchers analyzed mouse brain and embryonic stem cells, uncovering how spatial genome organization influences gene regulation.
- The technique provides the first simultaneous view of chromatin structure and transcriptional activity in individual cells.
- Open-access datasets and open-source code are publicly available, accelerating adoption across the research community.
Why Single-Cell Four-Omics Sequencing Represents a Paradigm Shift in Genomics
Traditional genomic methods analyze bulk populations of cells, averaging out critical variations between individual cells that drive development, disease progression, and response to treatments. Single-cell sequencing technologies have begun to address this limitation, but most approaches examine only one or two molecular layers at a time. The CHARM method breaks this barrier by integrating four complementary datasets from the same cell: 3D genome structure (conformation), histone modification patterns (epigenetic marks), chromatin accessibility (regulatory element activity), and gene expression (transcript levels). This multi-layered perspective enables researchers to directly connect genome folding with functional outcomes, revealing how structural changes in DNA influence which genes are expressed and when.
The Four Layers of Gene Regulation Now Accessible in Single Cells
The CHARM technique builds upon decades of research into how the genome operates within the nucleus. Genome conformation refers to the 3D folding of DNA, which brings distant regulatory elements like enhancers into physical proximity with gene promoters. Histone modifications are chemical tags added to histone proteins around which DNA is wound; these marks can either promote or repress gene expression depending on their type and location. Chromatin accessibility describes regions of the genome where DNA is loosely packaged and accessible to transcription machinery, indicating active regulatory elements. Finally, gene expression measures which genes are actively transcribed into RNA. By measuring all four simultaneously, scientists can establish causal relationships between genome structure, epigenetic state, and transcriptional output.
How the CHARM Method Works: Technical Breakthroughs Behind the Innovation
Developing CHARM required overcoming significant technical challenges, particularly in combining disparate data types from the same cell without losing resolution or introducing artifacts. The research team employed droplet-based microfluidics to isolate single nuclei while preserving their molecular contents. They then applied a series of highly optimized biochemical assays: Hi-C for genome conformation, CUT&Tag for histone modifications, ATAC-seq for chromatin accessibility, and Smart-seq for full-length transcriptomes. Crucially, the workflow was automated using 3D-printed liquid-handling robots to minimize human error and maximize throughput. All computational pipelines and data processing scripts are publicly available on GitHub, enabling other laboratories to replicate and extend the method.
Key Findings: New Insights into Genome Organization and Gene Control
The Nature study presents several groundbreaking findings that challenge and refine our understanding of genome biology. Most strikingly, the researchers discovered that regulatory elements—such as enhancers and promoters—tend to cluster together in specific spatial neighborhoods within the nucleus, forming what they term "regulatory hubs." These hubs persist across different cell types and developmental stages, suggesting they represent fundamental organizational units of the genome. The team also found that changes in genome architecture often precede changes in gene expression during development, indicating that 3D structure may act as a scaffold upon which epigenetic and transcriptional programs are built.
Dynamic Enhancer-Promoter Interactions Shape Cellular Identity
One of the most significant revelations involves the dynamic nature of enhancer-promoter interactions. Enhancers are DNA sequences that can activate gene expression from great distances by looping through 3D space to contact their target promoters. The CHARM data show that these loops are not static but constantly rearrange, with different enhancers competing for access to promoters depending on the cell's state. This competition helps explain how a single genome can give rise to hundreds of distinct cell types with specialized functions. For example, in neural stem cells, specific enhancer hubs coordinate the expression of genes required for neurogenesis, while repressing genes that would drive differentiation into other lineages.
Cell-Type-Specific Chromatin Landscapes Emerge from Multi-Omic Profiles
By analyzing thousands of single cells from mouse brain and embryonic stem cell cultures, the team mapped the chromatin landscape across diverse cell types. They identified cell-type-specific patterns of histone modifications that correlate with distinct gene expression programs. For instance, neurons showed high levels of H3K27ac—a mark of active enhancers—at genes involved in synaptic function, while stem cells displayed bivalent chromatin domains marked by both H3K27me3 (repressive) and H3K4me3 (active) at developmental genes, a hallmark of pluripotency. These patterns were invisible in bulk sequencing data, highlighting the importance of single-cell resolution.
Applications Across Biology: From Development to Disease
The implications of single-cell four-omics sequencing extend far beyond basic research. In developmental biology, the technique provides a window into how the genome transitions from a pluripotent state (capable of becoming any cell type) to specialized lineages. This is particularly relevant for understanding congenital disorders and regenerative medicine. In neuroscience, CHARM offers new ways to study how neural circuits form by mapping the regulatory landscape of different neuron types in the brain. For cancer research, the method could reveal how tumor cells hijack normal chromatin architecture to drive uncontrolled growth, potentially identifying new therapeutic targets.
Unraveling the Molecular Basis of Cognitive Traits and Disorders
The researchers also explored connections between chromatin organization and cognitive function by examining genes associated with intelligence in genome-wide association studies. They found that many of these genes, including SATB2 (a chromatin organizer linked to intellectual disability and schizophrenia), are located within regulatory hubs that coordinate their expression during brain development. This suggests that subtle disruptions in 3D genome architecture may underlie certain neurodevelopmental disorders. The CHARM dataset provides a resource for exploring these links in greater depth.
Open Science: Making Breakthrough Data and Tools Accessible
A distinguishing feature of this study is its commitment to open science. All raw and processed datasets have been deposited in public repositories, including the NCBI Sequence Read Archive (SRA) under BioProject accession PRJNA1284811 and the Gene Expression Omnibus (GEO) under accession GSE303006. The mouse reference genome assembly and annotations used are based on GRCm38 with GENCODE release M23, ensuring compatibility with existing genomic resources. Additionally, the team has made all computational code available on GitHub, along with 3D-printable models for the automated liquid-handling workflow. This open approach accelerates scientific progress by allowing researchers worldwide to build upon these findings without barriers.
Challenges and Future Directions: Scaling Up and Refining the Method
While CHARM represents a major advance, significant challenges remain. Current implementations require expensive equipment and specialized expertise, limiting widespread adoption. The throughput of single-cell multi-omics techniques is also lower than single-modality approaches, making large-scale studies resource-intensive. Additionally, integrating four distinct data types introduces computational complexity, requiring advanced machine learning algorithms to reconcile differences in scale and noise profiles across modalities. Future improvements may include integrating additional layers of regulation, such as DNA methylation or protein-DNA interactions, and developing more streamlined workflows to reduce cost and complexity.
Comparing CHARM to Other Single-Cell Multi-Omics Techniques
The CHARM method joins a rapidly expanding toolkit of single-cell multi-omics techniques, each with unique strengths and limitations. For example, scATAC-seq + RNA-seq (like the 10x Genomics multiome assay) profiles chromatin accessibility and transcription simultaneously but lacks information on 3D genome structure and histone modifications. Other methods like scCUT&Tag-seq or scTHS-seq focus on specific epigenetic marks but don't capture genome conformation. CHARM's advantage lies in its comprehensiveness, though this comes at the cost of higher technical complexity and lower throughput compared to simpler two-omics approaches.
The ability to measure four layers of regulation in the same cell transforms our understanding of gene control from a linear sequence of events into a dynamic, three-dimensional process where structure and function are inextricably linked. This work not only provides unprecedented insight into genome biology but also establishes a blueprint for future multi-omic studies across biology.
The Broader Impact: Shaping the Future of Personalized and Precision Medicine
As single-cell multi-omics techniques like CHARM mature, they are poised to revolutionize medicine by enabling precision diagnostics and therapies tailored to individual patients. For instance, in oncology, profiling tumor cells with four-omics sequencing could reveal how specific chromatin alterations drive drug resistance, guiding the selection of targeted therapies. In regenerative medicine, understanding the regulatory landscape of stem cells could improve protocols for generating replacement tissues. The insights gained from these techniques may also inform our understanding of aging, as changes in chromatin architecture are linked to cellular senescence and age-related diseases.
Expert Reactions: What Scientists Are Saying About the Breakthrough
Leading genomics researchers have praised the CHARM method for its technical rigor and scientific impact. Dr. Tony Kouzarides, a pioneer in chromatin biology at the University of Cambridge, noted that "integrating multiple layers of regulation in single cells provides a holistic view of genome function that was previously impossible. This work sets a new standard for epigenetic studies." Similarly, Dr. Bing Ren, director of the Center for Epigenomics at UCSD, commented that "the discovery of regulatory hubs and their dynamic nature opens exciting new avenues for understanding how genomes are organized and controlled."
How to Access and Use the CHARM Datasets and Tools
Researchers interested in exploring the CHARM datasets or replicating the method can access all resources through the provided repositories and code bases. The raw sequencing data are available in the NCBI SRA under BioProject PRJNA1284811, while processed datasets are deposited in GEO under accession GSE303006. For computational analysis, the team's GitHub repository (https://github.com/skelviper/CHARM) contains preprocessing scripts, data integration pipelines, and visualization tools. Additionally, the 3D-printable models for the automated liquid-handling workflow are available on a separate GitHub repository (https://github.com/skelviper/OT2), enabling labs to build cost-effective automation systems.
Conclusion: A New Era of Genome Biology
The development of single-cell four-omics sequencing marks a turning point in genomics, shifting the field from studying isolated layers of regulation to examining the genome as an integrated, dynamic system. The CHARM method provides a powerful new lens through which to view gene regulation, offering insights that will shape research across biology and medicine for decades to come. As the technology matures and becomes more accessible, it promises to unlock answers to some of the most profound questions about life, development, and disease. The future of genome science is not just multi-omic—it is multi-dimensional.
Frequently Asked Questions
- What is single-cell four-omics sequencing?
- Single-cell four-omics sequencing is a technique that simultaneously profiles four layers of gene regulation in individual cells: genome conformation (3D structure), histone modifications (epigenetic marks), chromatin accessibility (regulatory element activity), and gene expression (transcript levels). This multi-layered approach provides a comprehensive view of how genome structure and epigenetic modifications control gene activity.
- How does CHARM differ from other single-cell multi-omics methods?
- CHARM stands out by integrating four distinct data types in the same cell, whereas most other methods profile only two or three layers. For example, scATAC-seq + RNA-seq captures chromatin accessibility and transcription but lacks 3D genome structure and histone modification data. CHARM's comprehensiveness comes with higher technical complexity.
- What are the potential medical applications of this technology?
- The technology could revolutionize precision medicine by revealing how chromatin alterations drive diseases like cancer, neurological disorders, and developmental conditions. It may also improve regenerative medicine by clarifying the regulatory programs that maintain stem cell pluripotency or guide differentiation into specific cell types.